Autophagy Infection Immunity (A2I)
Centre International de Recherche en Infectiologie – CIRIINSERM U1111, CNRS UMR5308, ENS-Lyon, UCB-Lyon1
21 avenue Tony Garnier
69007 Lyon - Lyon
Site web - mathias.faure@inserm.fr -
Principal investigator
Pr Mathias Faure

Research themes
The team is interested in physiological and pathological aspects of human cellular defense mechanisms. To this end, the team has historically used as a biological model the measles virus which is an immunosuppressive virus. In 1993, the team of Prof. Chantal Rabourdin-Combe (1952-2011) identified the first human receptor for this virus, CD46. More recently, comprehensive studies related to the functions of this receptor has established the basis of the projects of the current team, putting autophagy at the center of our research projects. Thus, we have described a molecular link between CD46 and autophagy, which is engaged upon measles virus infection. We detailed then the close relationship between this virus and autophagy, describing for the first time that several waves of autophagy can be induced in the course of an infection. We also characterized the first interactome connecting many RNA viral proteins with dozens of key proteins involved in autophagy. This study highlighted the fact that common strategies could have been evolved by different viruses in order to manipulate autophagy. Recently we have shown the dual function of an autophagic receptor which, in addition to target pathogens towards the autophagic machinery, also regulates the maturation of autophagosome to efficiently degrade the substrates.
Currently, our research projects focus on three main axis :
- Regulation of Autophagy during infection
Based on interactions that we found between pathogen proteins and autophagy-associated proteins, we try to depict the molecular (de)regulation of autophagy in the course of infections. A focus concerns the stages of recognition of intracellular pathogens, their targeting to the autophagy machinery, and the regulation of the maturation of the pathogen-containing autophagosomes (their fusion with lysosomes). These studies will tell us on how autophagy per se is regulated at the molecular level and inform us about the pathways of possible diversion targeted by infectious agents. We also study the alterations of these interactions and their biological consequences in the context of co-infections, with pathogens which display opposite sensitivities towards autophagy.
- Autophagy and antigen presentation
Antigen presentation by molecules of the major histocompatibility complex (MHC) is an essential step to initiate an adaptive immune response to efficiently fight pathogens. Since numerous antigens which are presented by MHC molecules traffic via the lysosomes, autophagy is an important reservoir for the production of antigenic peptides derived from the degradation of intracellular pathogens. We seek to understand the molecular connections between autophagy and antigen presentation and to measure their impact on the diversity of the presentation, especially in the context of self antigens (those of the cell), and non-self antigens (those from pathogens).
- Autophagy and healthy/pathological human primary cells
Autophagy is particularly decrypted and analyzed using immortalized cell lines and appropriate animal models. However, few studies focus on the analysis of primary cells, although in humans, many diseases are associated with polymorphisms of autophagic genes. We seek to standardize the study of autophagy in immune cells from healthy donors to better understand autophagy failures under pathological conditions. We pay a special attention to inflammatory bowel diseases, in particular Crohn’s disease for which many genes of susceptibility are linked to autophagy.
Descriptive figure
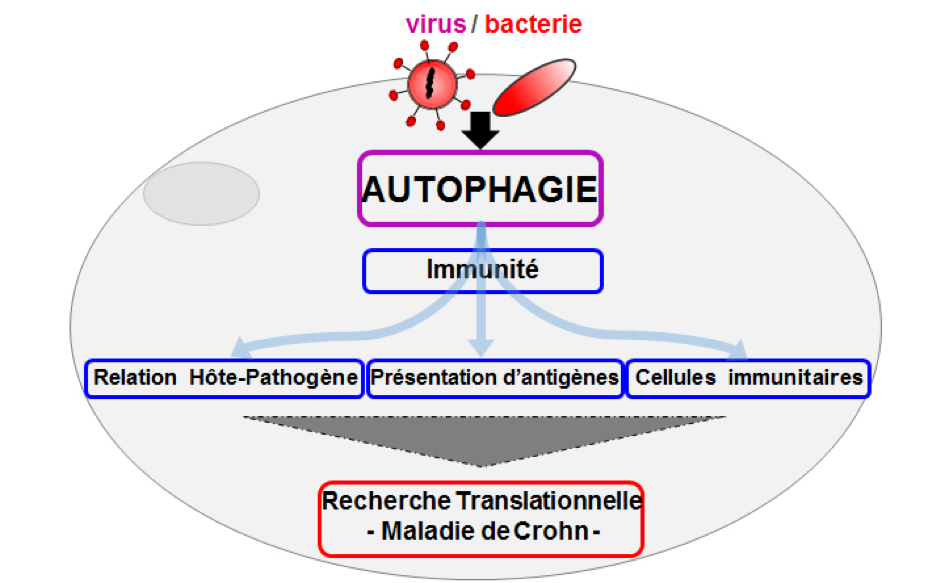
Publications
– C. Viret and M. Faure (2015). Autophagy in immune responses to viruses. Book title: Autophagy, Infection, and the Immune Response , Wiley Blackwell press edition, USA. p279-302.
– Verlhac P., Viret C. and Faure M (2015). Dual Function of CALCOCO2/NDP52 during Xenophagy. Autophagy V11 p965-6 (puncta).
– Verlhac P., Viret C and Faure M (2015). NDP52, Autophagy and Pathogens : “The war then ceased through lack of combatants”. Med/Sci (Paris) V6-7 p594-7.
– M. Faure. (2014). The p-value of HPIV3-mediated autophagy inhibition. Cell Host & Microbe V15 p519-521 (preview).
– C. Richetta, I. P. Grégoire, P. Verlhac, O. Azocar, J. Baguet, M. Flacher, F. Tangy, C. Rabourdin-Combe and M. Faure. (2013). Sustained Autophagy Contributes to Measles Virus Infectivity. PloS Pathogens, Sept;9(9):e1003599.
– M. Faure and F. Lafont. (2013). Pathogen-induced autophagy signaling in innate immunity. J Innate Immunol., V5 p456-470 (review).
– Petkova DS, Viret C, M. Faure (2013). IRGM in autophagy and viral infections. Frontiers in Immunology, V3 Art.426 (review).
– C. Richetta and M. Faure. (2013). Autophagy in innate antiviral immunity. Cell Microbiol., V15 p368-376 (review).
– I. Pombo Grégoire, C. Rabourdin-Combe and M. Faure (2012). Autophagy and RNA virus interactomes reveal IRGM as a common target. Autophagy, V8 p1136-1137 (puncta)
– P.-E. Joubert, I. Pombo Gregoire, G. Meiffren, C. Rabourdin-Combe, and M. Faure (2011). Autophagy and Pathogens : Bon appetit !. Med/Sci (Paris) V27 p41-47.
– I. Pombo Grégoire, C. Richetta, L. Meyniel-Schicklin, S. Borel, F. Pradezynski, O. Diaz, A. Deloire, O. Azocar, J. Baguet, M. Le Breton, P. E. Mangeot, V. Navratil, P.-E. Joubert, M. Flacher, P.-O. Vidalain, P. André, V. Lotteau, M. Biard-Piechaczyk, C. Rabourdin-Combe* and M. Faure* (2011). IRGM is a common target of RNA viruses that subvert the Autophagy network. PloS Pathogens, december, V7 : e1002422.
– G. Meiffren, P.-E. Joubert, I. Pombo Gregoire, P. Codogno, C. Rabourdin-Combe, M. Faure (2010). Pathogen recognition by the cell surface receptor CD46 induces autophagy. Autophagy, V6 p299-300 (puncta).
– Joubert, PE, Meiffren, G, Pombo Gregoire, I, Pontini, G, , Richetta, C, Flacher, M, Azocar, O, Vidalain, PO, Vidal, M, Lotteau, V, Codogno, P, Rabourdin-Combe, C and Faure, M (2009). Autophagy Induction by the Pathogen Receptor CD46. Cell Host & Microbe, V6 p354-366.
Composition de l'équipe
Christophe Viret (CR CNRS)
Aurore Rozières (MCF Université Lyon 1)
Stéphane Nancey (PU-PH Lyon Sud)
Bernard Flourié (PU-PH Lyon Sud)
Joël Baguet (IR INSERM)
Cécile Daussy-Abes (AI INSERM)
Pauline Verlhac (doctorante)
Mathieu Claviere (doctorant)
Gilles Boschetti (PHU Lyon Sud)
Emilien Gimaret (M2-Recherche Lyon 1)
Céline Berquet (M1-EPHE)